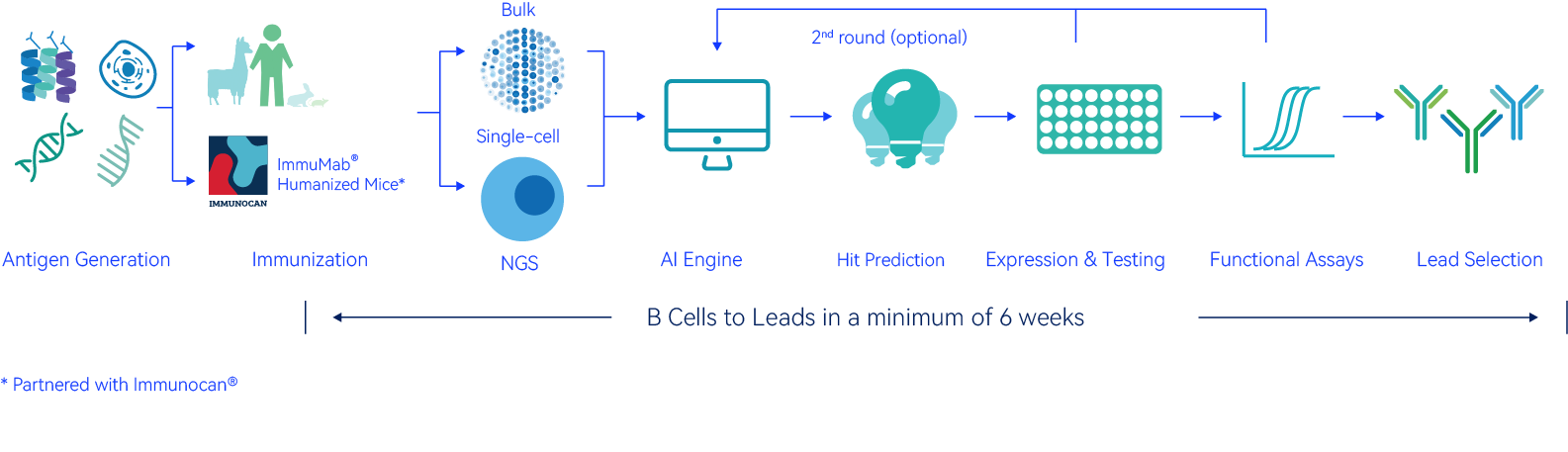
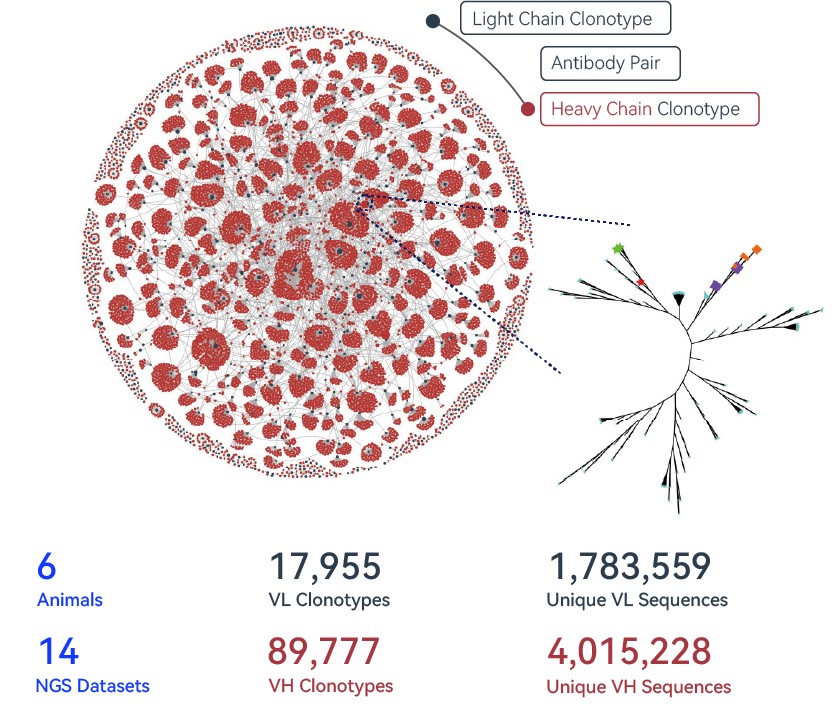
The immunized BCR repertoire is huge and contains numerous sequences with therapeutic potential.
XploreSeqTM screens through this repertoire to identify hits, powered by our AI engine that has been trained and refined over the years. A typical discovery program could generate millions of BCR sequences (shown above) and poses tremendous challenges for finding the needle in the haystack.
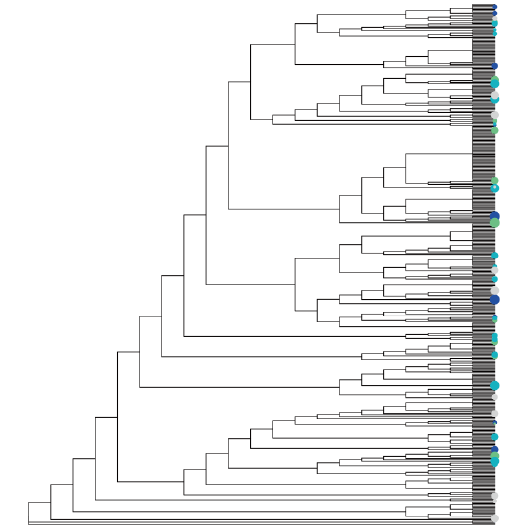
Reconstructed antibody lineage tree. Visualization from real program.
We believe in both the quality & diversity of candidates. The objective is to pick antibodies from different lineages with high confidence of target-binding and excellent developa-bility right out of the box. This gives us a diverse panel of developable hits that could cover a maximum number of epitopes, leading to higher success rate in downstream assays.
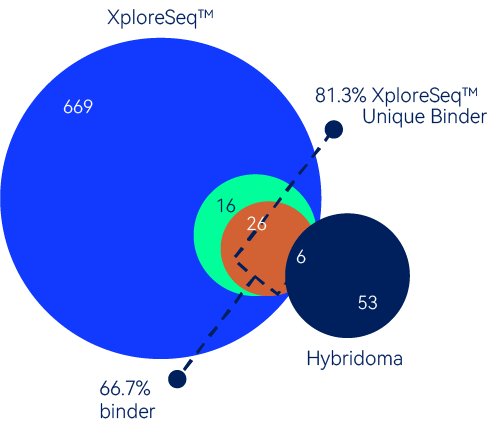
For this campaign, XploreSeq™ predicted 717 binders, from which 48 were randomly chosen for expression and testing. 66.7% (32/48) were confirmed to be binders, and only 18.6% (6/32) binders overlap with hybridoma hits.
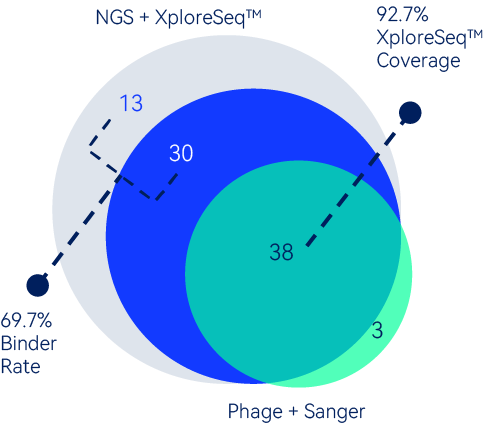
For this campaign, a total of 41 unique VHH hits were identified by immune phage library through Sanger sequencing.
The library was then NGS sequenced and subject to XploreSeqTManalysis. NGS results include 38 of the 41 hits identified previously.
In addition, XploreSeqTM predicted 43 new VHH binders. When expressed and tested, 30 of these 43 (69.7%) were confirmed to be binders, which significantly expanded the diversity of hits obtained by phage display alone.
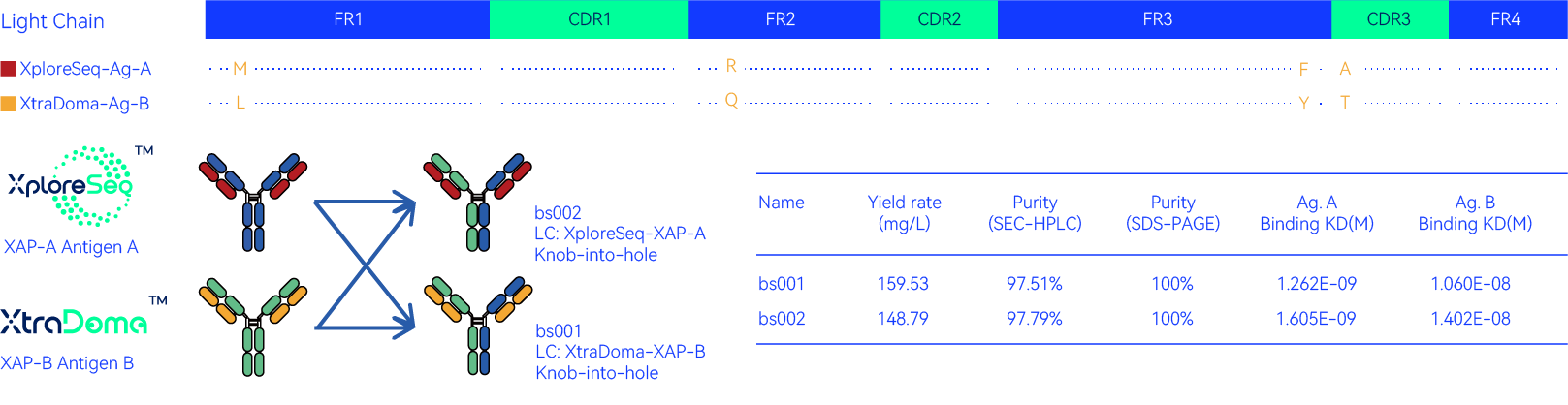
Light chains exhibit less diversity in immune repertoires, which facilitates the identification of common light chains across different repertoires for making bispecific antibodies. XploreSeqTM is uniquely positioned for a such task, as it collects light chain sequences in the millions, greatly amplifying the likelihood of finding a common LC.
In this case, XploreSeqTM was able to identify a similar light chain in antigen A repertoire with a hit identified by our Xtra-DomaTM hybridoma platform for antigen B. We directly built two common LC bispecific antibodies based on both of these LCs. Both molecules expressed well, showed high purity and bound both targets as intended.
XtalPi is keenly aware that every research partner has unique needs and priorities, and is happy to create a customized solution to optimize your research program





